This vignette covers the multivariate normal (MVN), affine
transformations, and mixture distribution facilities in
algebraic.dist. These features were introduced in version
0.3.0 and provide closed-form operations wherever possible, falling back
to Monte Carlo only when necessary.
MVN Basics
Construct an MVN with mvn(). The mu
argument is the mean vector and sigma is the
variance-covariance matrix (defaulting to the identity).
M <- mvn(mu = c(1, 2, 3),
sigma = matrix(c(1.0, 0.5, 0.2,
0.5, 2.0, 0.3,
0.2, 0.3, 1.5), nrow = 3, byrow = TRUE))
M
#> Multivariate normal distribution (3 dimensions)
#> mu:
#> [1] 1 2 3
#> sigma:
#> [,1] [,2] [,3]
#> [1,] 1.0 0.5 0.2
#> [2,] 0.5 2.0 0.3
#> [3,] 0.2 0.3 1.5Standard accessors return the moments and dimensionality.
mean(M)
#> [1] 1 2 3
vcov(M)
#> [,1] [,2] [,3]
#> [1,] 1.0 0.5 0.2
#> [2,] 0.5 2.0 0.3
#> [3,] 0.2 0.3 1.5
dim(M)
#> [1] 3The params() generic flattens all parameters into a
single named vector (useful for optimisation interfaces).
params(M)
#> mu1 mu2 mu3 sigma1 sigma2 sigma3 sigma4 sigma5 sigma6 sigma7 sigma8
#> 1.0 2.0 3.0 1.0 0.5 0.2 0.5 2.0 0.3 0.2 0.3
#> sigma9
#> 1.5Sampling
sampler() returns a reusable sampling function. The
output is an n-by-d matrix where rows are observations and columns are
dimensions.
Marginals
Extracting a marginal over a subset of indices returns a new MVN (or
a univariate normal when a single index is selected).
M13 <- marginal(M, c(1, 3))
M13
#> Multivariate normal distribution (2 dimensions)
#> mu:
#> [1] 1 3
#> sigma:
#> [,1] [,2]
#> [1,] 1.0 0.2
#> [2,] 0.2 1.5
mean(M13)
#> [1] 1 3
vcov(M13)
#> [,1] [,2]
#> [1,] 1.0 0.2
#> [2,] 0.2 1.5Selecting a single component yields a univariate normal.
Closed-Form MVN Conditioning
Conditioning a multivariate normal on observed values uses the Schur complement formula. Given an MVN partitioned into free variables and observed variables , the conditional distribution is:
This is exact – no Monte Carlo is involved.
M2 <- mvn(mu = c(0, 0), sigma = matrix(c(1, 0.8, 0.8, 1), 2, 2))
# Condition on X2 = 1
M_cond <- conditional(M2, given_indices = 2, given_values = 1)
mean(M_cond)
#> [1] 0.8
vcov(M_cond)
#> [1] 0.36With , the conditional mean is , and the conditional variance is .
Predicate-based MC fallback
When the conditioning event is not “observe variable
at value
”
but an arbitrary predicate, the package falls back to Monte Carlo. This
is accessed through the P argument.
set.seed(42)
M_cond_mc <- conditional(M2, P = function(x) abs(x[2] - 1) < 0.1)
mean(M_cond_mc)
#> [1] 0.7938476 0.9969783The MC result is approximate and depends on the tolerance in the predicate. Prefer the closed-form interface whenever possible.
Affine Transformations
affine_transform(x, A, b) computes the distribution of
where
.
The result is:
This is a closed-form operation that returns a new MVN (or
normal if the result is one-dimensional).
Portfolio example
Consider two assets with expected returns and a covariance structure. A portfolio is a linear combination of the returns.
returns <- mvn(
mu = c(0.05, 0.08),
sigma = matrix(c(0.04, 0.01,
0.01, 0.09), 2, 2)
)
# 60/40 portfolio weights
w <- matrix(c(0.6, 0.4), nrow = 1)
portfolio <- affine_transform(returns, A = w)
mean(portfolio)
#> [1] 0.062
vcov(portfolio)
#> [1] 0.0336The portfolio return is . The portfolio variance is , a scalar that captures the diversification benefit.
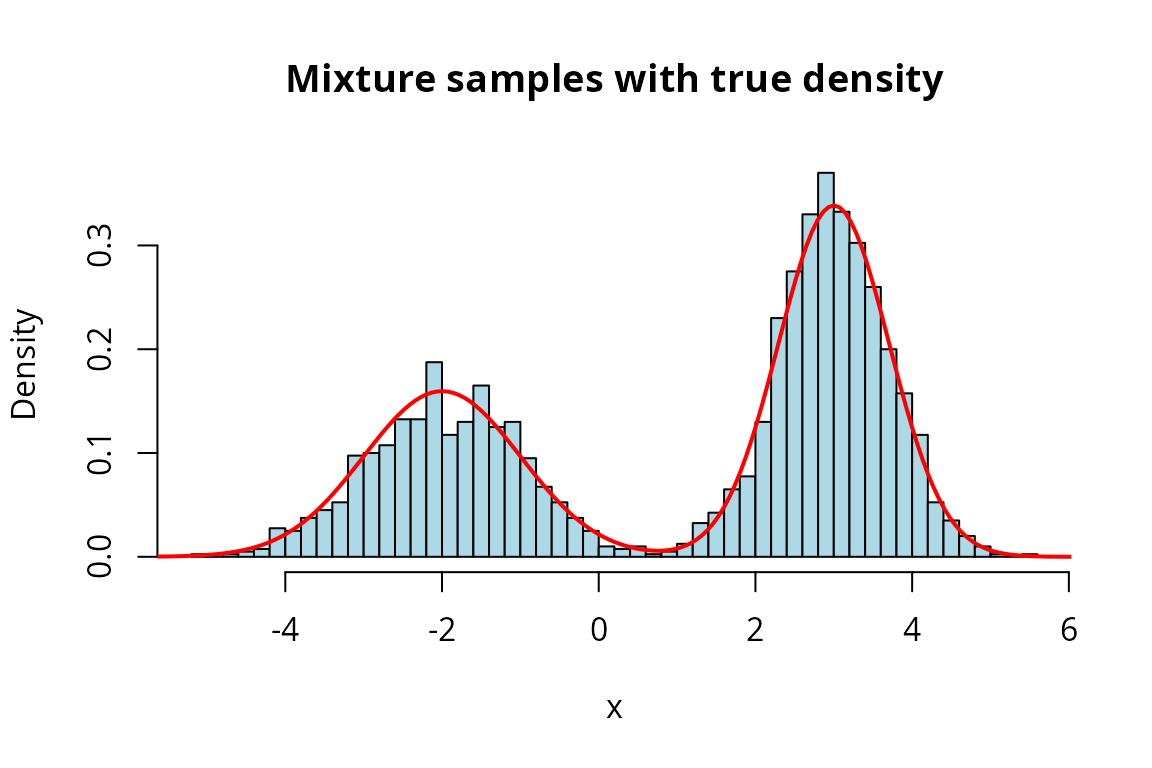
Mixture Distribution Basics
A mixture distribution is constructed from a list of component distributions and a vector of non-negative weights that sum to one.
m <- mixture(
components = list(normal(-2, 1), normal(3, 0.5)),
weights = c(0.4, 0.6)
)
m
#> Mixture distribution (2 components)
#> [w=0.400] Normal distribution (mu = -2, var = 1)
#> [w=0.600] Normal distribution (mu = 3, var = 0.5)Mean and variance
The mixture mean is the weighted average of component means: .
mean(m)
#> [1] 1The mixture variance uses the law of total variance: .
vcov(m)
#> [1] 6.7The between-component term captures the spread of the component means around the overall mean. This is why mixtures can have much larger variance than any individual component.
Density and CDF
density() and cdf() return callable
functions, following the same pattern as other distributions in the
package.
f <- density(m)
x_vals <- seq(-6, 8, length.out = 200)
y_vals <- sapply(x_vals, f)
plot(x_vals, y_vals, type = "l", lwd = 2,
main = "Bimodal mixture density",
xlab = "x", ylab = "f(x)")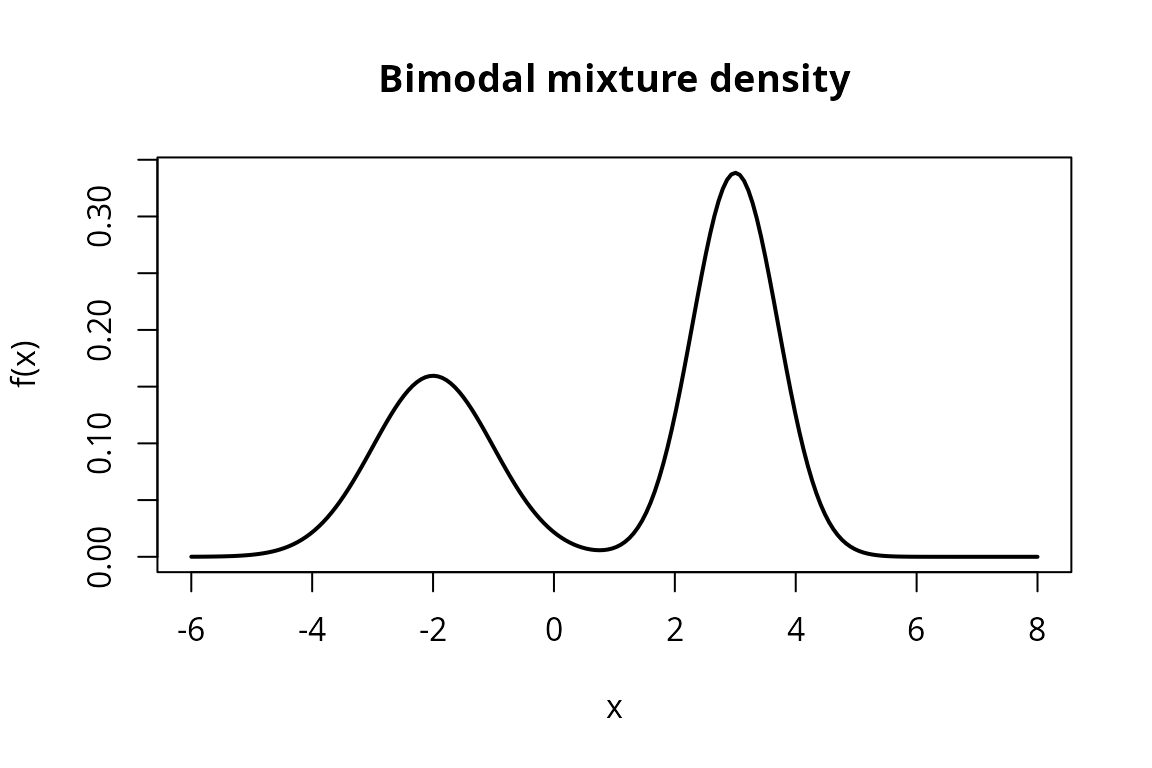
The bimodal shape is clearly visible, with peaks near and .
F <- cdf(m)
F_vals <- sapply(x_vals, F)
plot(x_vals, F_vals, type = "l", lwd = 2,
main = "Mixture CDF",
xlab = "x", ylab = "F(x)")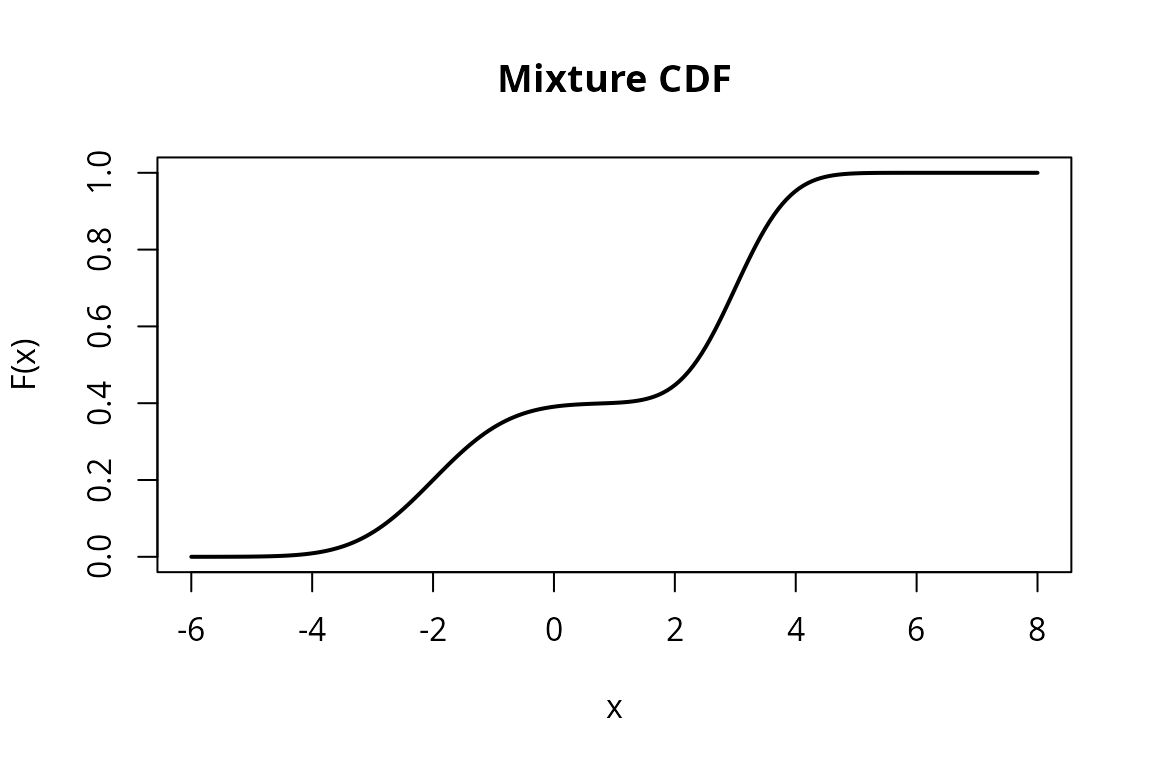
Mixture Operations
Marginals of mixtures
The marginal of a mixture is a mixture of marginals with the same weights. This follows directly from the definition: .
m2 <- mixture(
list(mvn(c(0, 0), diag(2)), mvn(c(3, 3), diag(2))),
c(0.5, 0.5)
)
m2_x1 <- marginal(m2, 1)
m2_x1
#> Mixture distribution (2 components)
#> [w=0.500] Normal distribution (mu = 0, var = 1)
#> [w=0.500] Normal distribution (mu = 3, var = 1)
mean(m2_x1)
#> [1] 1.5The marginal mean is , the weighted average of the component marginal means.
Conditional of mixtures (Bayesian weight update)
Conditioning a mixture on observed values updates the component weights via Bayes’ rule. Each component’s weight is scaled by its marginal density at the observed value:
Components whose marginal density is higher at the observed value receive more weight.
mc <- conditional(m2, given_indices = 2, given_values = 0)
mc
#> Mixture distribution (2 components)
#> [w=0.989] Normal distribution (mu = 0, var = 1)
#> [w=0.011] Normal distribution (mu = 3, var = 1)
mean(mc)
#> [1] 0.03296083Conditioning on shifts weight toward the first component (whose mean for is 0) and away from the second component (whose mean for is 3). The conditional mean of is therefore pulled closer to 0 than the unconditional value of 1.5.
We can also condition at a value near the second component to see the opposite effect.
mc_high <- conditional(m2, given_indices = 2, given_values = 3)
mc_high
#> Mixture distribution (2 components)
#> [w=0.011] Normal distribution (mu = 0, var = 1)
#> [w=0.989] Normal distribution (mu = 3, var = 1)
mean(mc_high)
#> [1] 2.967039Now the weight shifts toward the second component, and the conditional mean of is pulled toward 3.
Practical Example: Gaussian Mixture Classification
Gaussian mixture models (GMMs) are widely used for soft classification. The idea: each component represents a class, and conditioning on observed features updates the class probabilities.
Setup: a two-class GMM
Consider two clusters in 2D, representing two classes with equal prior probability.
class_a <- mvn(c(-2, -1), matrix(c(1.0, 0.3, 0.3, 0.8), 2, 2))
class_b <- mvn(c(2, 1.5), matrix(c(0.6, -0.2, -0.2, 1.2), 2, 2))
gmm <- mixture(
components = list(class_a, class_b),
weights = c(0.5, 0.5)
)
gmm
#> Mixture distribution (2 components)
#> [w=0.500] Multivariate normal distribution (2 dimensions)
#> [w=0.500] Multivariate normal distribution (2 dimensions)Visualise the clusters
set.seed(314)
samp_a <- sampler(class_a)(300)
samp_b <- sampler(class_b)(300)
plot(samp_a[, 1], samp_a[, 2], col = "steelblue", pch = 16, cex = 0.6,
xlim = c(-6, 6), ylim = c(-5, 5),
xlab = expression(X[1]), ylab = expression(X[2]),
main = "Two-class GMM samples")
points(samp_b[, 1], samp_b[, 2], col = "tomato", pch = 16, cex = 0.6)
legend("topright", legend = c("Class A", "Class B"),
col = c("steelblue", "tomato"), pch = 16)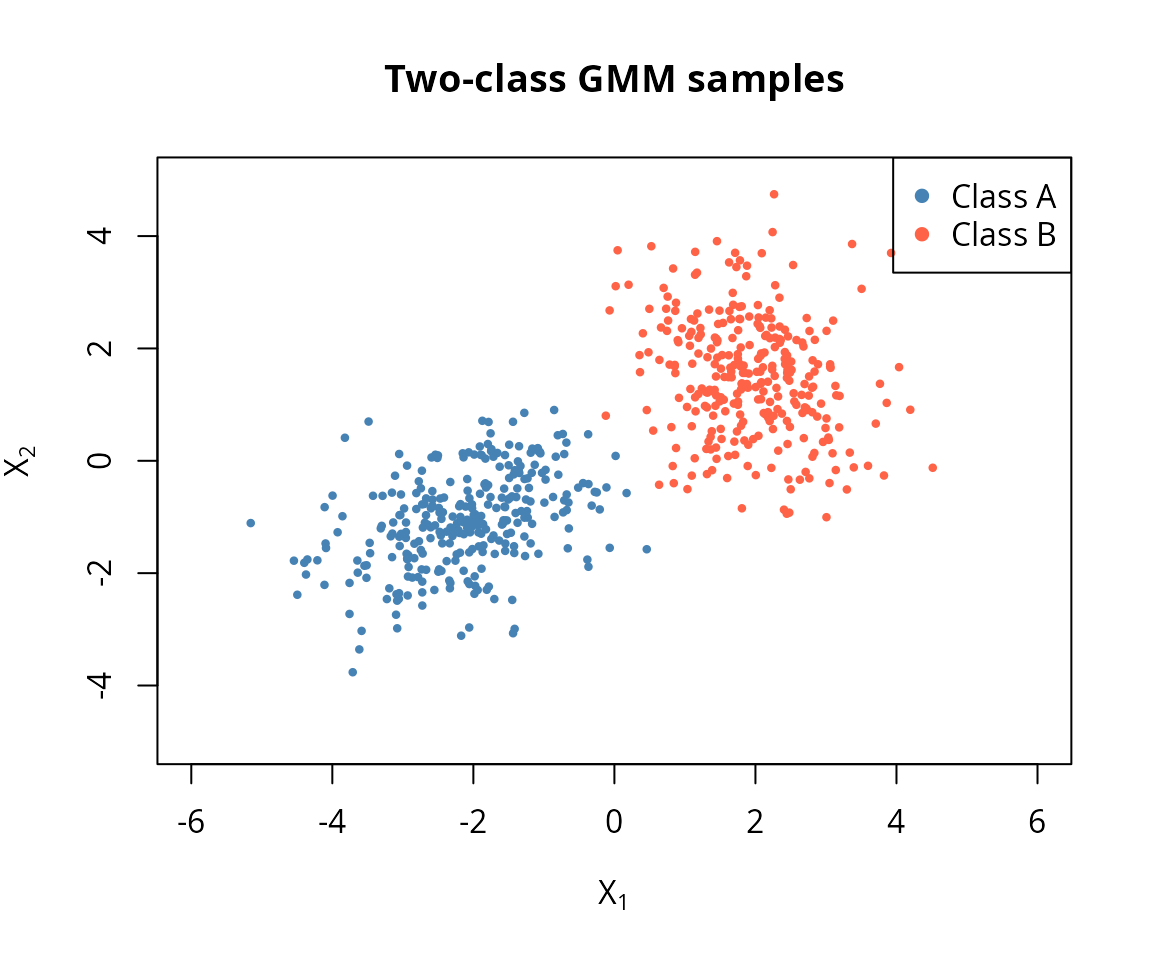
Soft classification by conditioning
Suppose we observe . Conditioning updates the class weights based on how likely each component is to produce that value.
post_neg <- conditional(gmm, given_indices = 1, given_values = -1)
post_neg
#> Mixture distribution (2 components)
#> [w=0.999] Normal distribution (mu = -0.7, var = 0.71)
#> [w=0.001] Normal distribution (mu = 2.5, var = 1.13333)The updated weights give the posterior class probabilities. Since is much more likely under Class A (centred at ) than Class B (centred at ), Class A receives most of the weight.
The conditional distribution of given is itself a mixture of the two conditional normals.
f_post <- density(post_neg)
x2_grid <- seq(-5, 5, length.out = 200)
y_post <- sapply(x2_grid, f_post)
plot(x2_grid, y_post, type = "l", lwd = 2,
xlab = expression(X[2]),
ylab = expression(f(X[2] ~ "|" ~ X[1] == -1)),
main = "Posterior density of X2 given X1 = -1")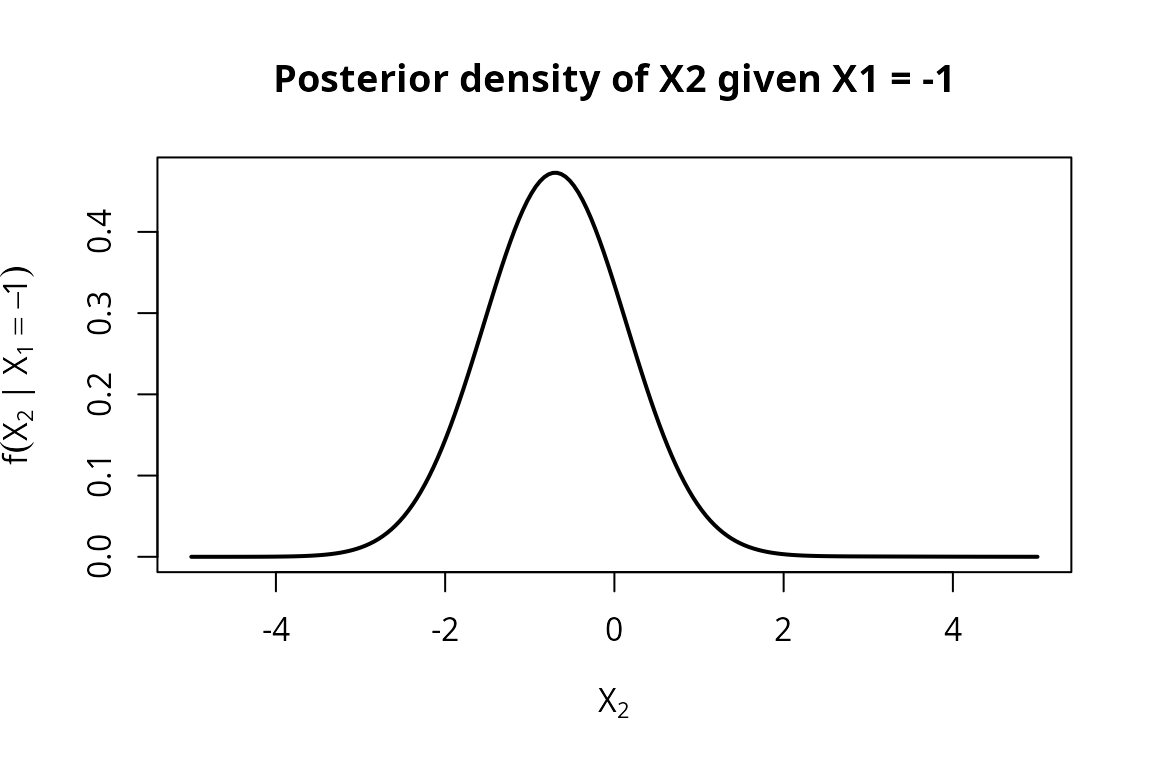
Now observe , which sits near the centre of Class B.
post_pos <- conditional(gmm, given_indices = 1, given_values = 2)
post_pos
#> Mixture distribution (2 components)
#> [w=0.000] Normal distribution (mu = 0.2, var = 0.71)
#> [w=1.000] Normal distribution (mu = 1.5, var = 1.13333)As expected, the weight shifts heavily toward Class B. The conditional density of now concentrates around the Class B conditional mean.
f_post_pos <- density(post_pos)
y_post_pos <- sapply(x2_grid, f_post_pos)
plot(x2_grid, y_post_pos, type = "l", lwd = 2,
xlab = expression(X[2]),
ylab = expression(f(X[2] ~ "|" ~ X[1] == 2)),
main = "Posterior density of X2 given X1 = 2")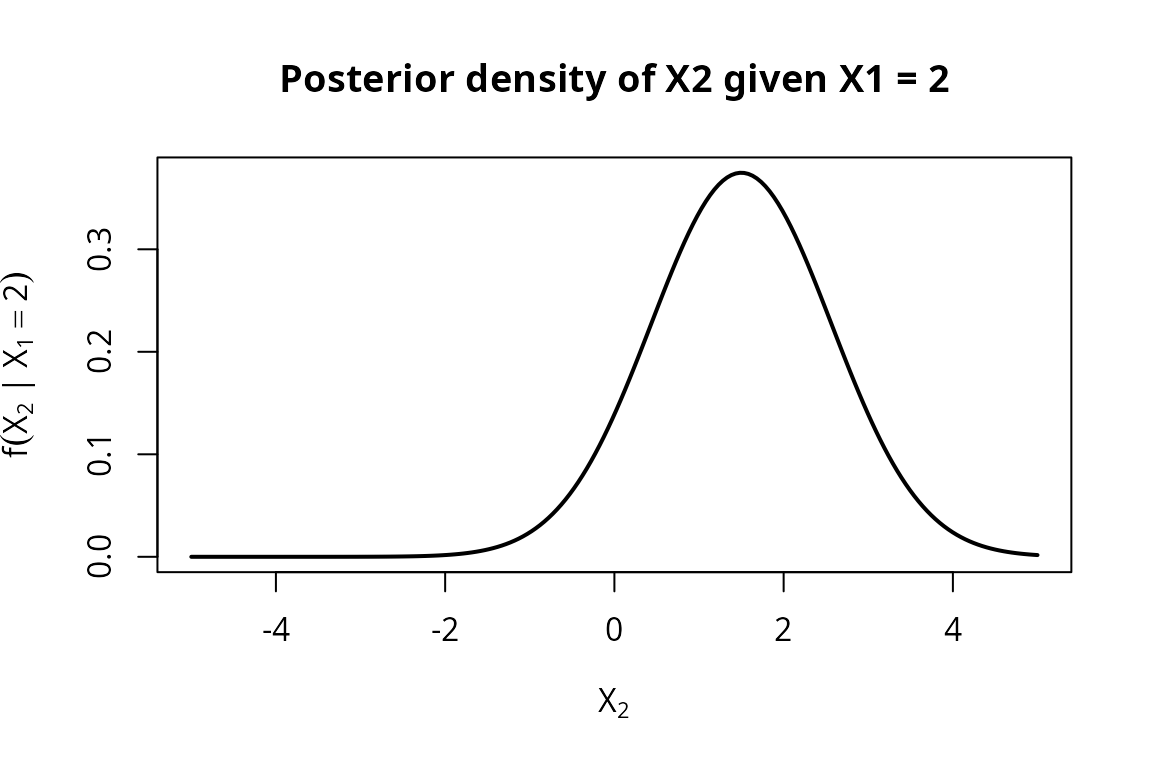
Ambiguous observations
When the observation falls between the two cluster means, both classes retain meaningful probability and the posterior becomes bimodal.
post_mid <- conditional(gmm, given_indices = 1, given_values = 0)
post_mid
#> Mixture distribution (2 components)
#> [w=0.746] Normal distribution (mu = -0.4, var = 0.71)
#> [w=0.254] Normal distribution (mu = 2.16667, var = 1.13333)
f_post_mid <- density(post_mid)
y_post_mid <- sapply(x2_grid, f_post_mid)
plot(x2_grid, y_post_mid, type = "l", lwd = 2,
xlab = expression(X[2]),
ylab = expression(f(X[2] ~ "|" ~ X[1] == 0)),
main = "Posterior density of X2 given X1 = 0 (ambiguous)")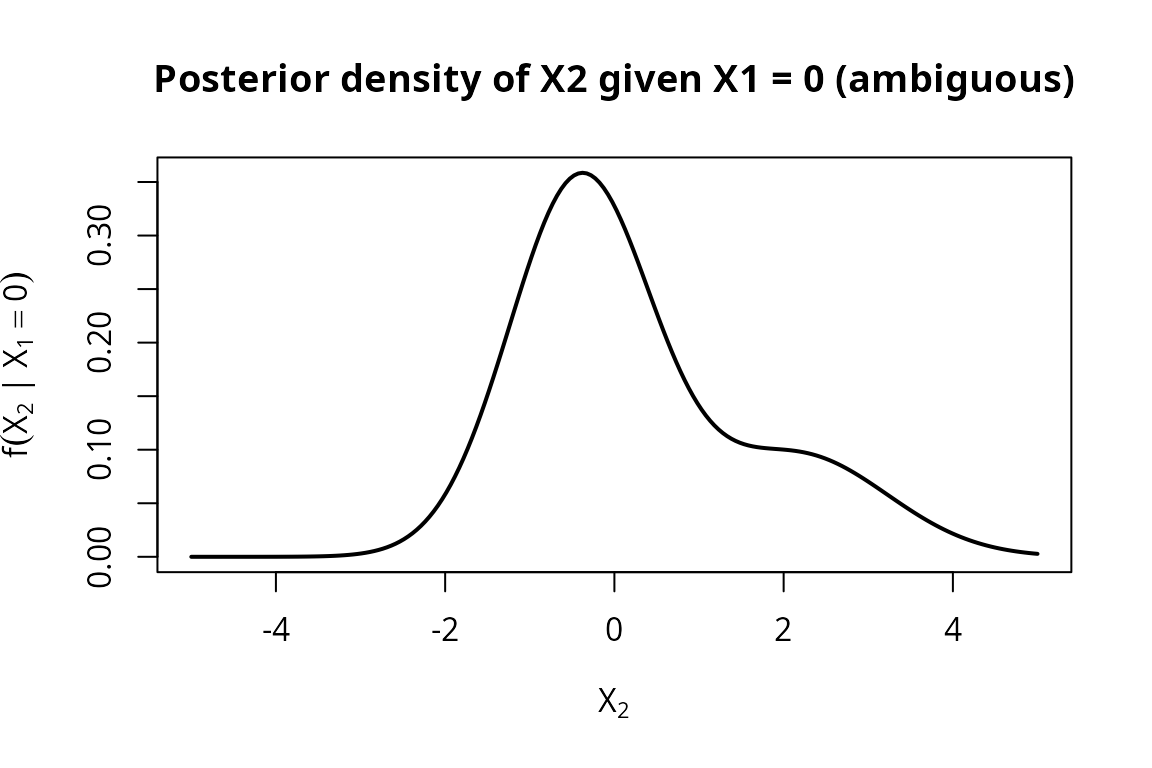
At the posterior for is bimodal – one mode near the Class A conditional mean, one near Class B. Although is equidistant from the two class means ( and ), Class A receives roughly 75% of the posterior weight because its broader variance ( vs ) gives it higher density at the midpoint. Class B still retains about 25% of the weight, so neither class is overwhelmingly dominant. This is the hallmark of soft classification: rather than forcing a hard decision, the mixture posterior represents genuine uncertainty about the class label.